This example has been auto-generated from the examples/ folder at GitHub repository.
Hierarchical Gaussian Filter
# Activate local environment, see `Project.toml`
import Pkg; Pkg.activate(".."); Pkg.instantiate();In this demo the goal is to perform approximate variational Bayesian Inference for Univariate Hierarchical Gaussian Filter (HGF).
Simple HGF model can be defined as:
\[\begin{aligned} x^{(j)}_k & \sim \, \mathcal{N}(x^{(j)}_{k - 1}, f_k(x^{(j - 1)}_k)) \\ y_k & \sim \, \mathcal{N}(x^{(j)}_k, \tau_k) \end{aligned}\]
where $j$ is an index of layer in hierarchy, $k$ is a time step and $f_k$ is a variance activation function. RxInfer.jl export Gaussian Controlled Variance (GCV) node with $f_k = \exp(\kappa x + \omega)$ variance activation function. By default the node uses Gauss-Hermite cubature with a prespecified number of approximation points in the cubature. In this demo we also show how we can change the hyperparameters in different approximation methods (iin this case Gauss-Hermite cubature) with the help of metadata structures. Here how our model will look like with the GCV node:
\[\begin{aligned} z_k & \sim \, \mathcal{N}(z_{k - 1}, \mathcal{\tau_z}) \\ x_k & \sim \, \mathcal{N}(x_{k - 1}, \exp(\kappa z_k + \omega)) \\ y_k & \sim \, \mathcal{N}(x_k, \mathcal{\tau_y}) \end{aligned}\]
In this experiment we will create a single time step of the graph and perform variational message passing filtering alrogithm to estimate hidden states of the system. For a more rigorous introduction to Hierarchical Gaussian Filter we refer to Ismail Senoz, Online Message Passing-based Inference in the Hierarchical Gaussian Filter paper.
For simplicity we will consider $\tau_z$, $\tau_y$, $\kappa$ and $\omega$ known and fixed, but there are no principled limitations to make them random variables too.
To model this process in RxInfer, first, we start with importing all needed packages:
using RxInfer, BenchmarkTools, Random, Plots, StableRNGsNext step, is to generate some synthetic data:
function generate_data(rng, n, k, w, zv, yv)
z_prev = 0.0
x_prev = 0.0
z = Vector{Float64}(undef, n)
v = Vector{Float64}(undef, n)
x = Vector{Float64}(undef, n)
y = Vector{Float64}(undef, n)
for i in 1:n
z[i] = rand(rng, Normal(z_prev, sqrt(zv)))
v[i] = exp(k * z[i] + w)
x[i] = rand(rng, Normal(x_prev, sqrt(v[i])))
y[i] = rand(rng, Normal(x[i], sqrt(yv)))
z_prev = z[i]
x_prev = x[i]
end
return z, x, y
endgenerate_data (generic function with 1 method)# Seed for reproducibility
seed = 42
rng = StableRNG(seed)
# Parameters of HGF process
real_k = 1.0
real_w = 0.0
z_variance = abs2(0.2)
y_variance = abs2(0.1)
# Number of observations
n = 300
z, x, y = generate_data(rng, n, real_k, real_w, z_variance, y_variance);Let's plot our synthetic dataset. Lines represent our hidden states we want to estimate using noisy observations.
let
pz = plot(title = "Hidden States Z")
px = plot(title = "Hidden States X")
plot!(pz, 1:n, z, label = "z_i", color = :orange)
plot!(px, 1:n, x, label = "x_i", color = :green)
scatter!(px, 1:n, y, label = "y_i", color = :red, ms = 2, alpha = 0.2)
plot(pz, px, layout = @layout([ a; b ]))
end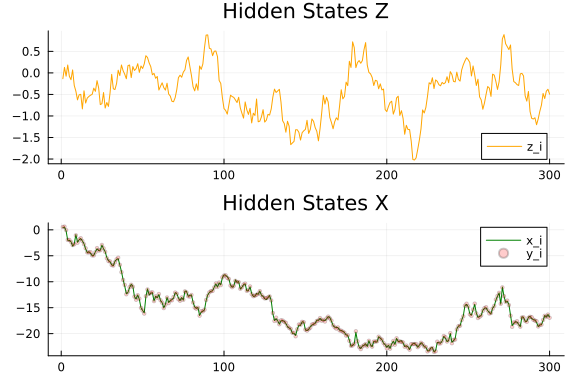
To create a model we use the @model macro:
# We create a single-time step of corresponding state-space process to
# perform online learning (filtering)
@model function hgf(y, κ, ω, z_variance, y_variance, z_prev_mean, z_prev_var, x_prev_mean, x_prev_var)
z_prev ~ Normal(mean = z_prev_mean, variance = z_prev_var)
x_prev ~ Normal(mean = x_prev_mean, variance = x_prev_var)
# Higher layer is modelled as a random walk
z_next ~ Normal(mean = z_prev, variance = z_variance)
# Lower layer is modelled with `GCV` node
x_next ~ GCV(x_prev, z_next, κ, ω)
# Noisy observations
y ~ Normal(mean = x_next, variance = y_variance)
end
@constraints function hgfconstraints()
# Structuted factorization constraints
q(x_next, x_prev, z_next) = q(x_next)q(x_prev)q(z_next)
end
@meta function hgfmeta()
# Lets use 31 approximation points in the Gauss Hermite cubature approximation method
GCV() -> GCVMetadata(GaussHermiteCubature(31))
endhgfmeta (generic function with 1 method)The code below uses the infer function from RxInfer to generate the message passing algorithm given the model and constraints specification. We also specify the @autoupdates in order to set new priors for the next observation based on posteriors.
function run_inference(data, real_k, real_w, z_variance, y_variance)
autoupdates = @autoupdates begin
# The posterior becomes the prior for the next time step
z_prev_mean, z_prev_var = mean_var(q(z_next))
x_prev_mean, x_prev_var = mean_var(q(x_next))
end
init = @initialization begin
q(x_next) = NormalMeanVariance(0.0, 5.0)
q(z_next) = NormalMeanVariance(0.0, 5.0)
end
return infer(
model = hgf(κ = real_k, ω = real_w, z_variance = z_variance, y_variance = y_variance),
constraints = hgfconstraints(),
meta = hgfmeta(),
data = (y = data, ),
autoupdates = autoupdates,
keephistory = length(data),
historyvars = (
x_next = KeepLast(),
z_next = KeepLast()
),
initialization = init,
iterations = 5,
free_energy = true,
)
endrun_inference (generic function with 1 method)Everything is ready to run the algorithm. We used the online version of the algorithm, thus we need to fetch the history of the posterior estimation instead of the actual posteriors.
result = run_inference(y, real_k, real_w, z_variance, y_variance);
mz = result.history[:z_next];
mx = result.history[:x_next];let
pz = plot(title = "Hidden States Z")
px = plot(title = "Hidden States X")
plot!(pz, 1:n, z, label = "z_i", color = :orange)
plot!(pz, 1:n, mean.(mz), ribbon = std.(mz), label = "estimated z_i", color = :teal)
plot!(px, 1:n, x, label = "x_i", color = :green)
plot!(px, 1:n, mean.(mx), ribbon = std.(mx), label = "estimated x_i", color = :violet)
plot(pz, px, layout = @layout([ a; b ]))
end
As we can see from our plot, estimated signal resembles closely to the real hidden states with small variance. We maybe also interested in the values for Bethe Free Energy functional:
plot(result.free_energy_history, label = "Bethe Free Energy")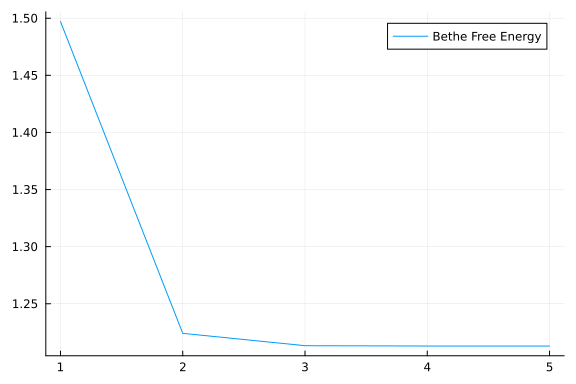
As we can see BetheFreeEnergy converges nicely to a stable point.